IDEAYA is pursuing precision medicine therapeutics, with a broad pipeline based on synthetic lethality – an emerging area of precision medicine oncology. Darovasertib (IDE196), a PKC inhibitor, is a clinical stage development compound for patients with tumors harboring GNAQ or GNA11 mutations, in ocular melanoma, including metastatic uveal melanoma (MUM) and primary uveal melanoma (UM). IDE397, a MAT2A inhibitor, is a clinical stage development compound for patients with tumors having MTAP gene deletion, estimated to represent approximately 15% of all solid tumors. IDE161, a PARG inhibitor, is a clinical-stage development candidate for patients having tumors with homologous recombination deficiency (HRD), including ER+, Her2-, HRD breast cancer and HRD ovarian cancer.

We are actively pursuing the discovery and development of precision medicine therapeutics for oncogenic and synthetic lethality targets in solid tumors. Our synthetic lethality programs target defined patient populations with significant prevalence, and as such, have the potential to be broadly impactful.
Our pipeline includes programs targeting:
- PKC, in tumor cells having activating mutations in GNAQ / GNA11 or , which occurs in greater than 90% of uveal melanoma;
- MAT2A, in tumor cells having MTAP gene deletion, which occurs in approximately 15% of all solid tumors;
- PARG, in tumors with HRD (e.g., BRCA) mutations or tumors having replication stress gene signature, including in ER+, Her2-, HRD breast cancer tumors, which reflects approximately 10-14% of breast cancer, and HRD ovarian cancer tumors, representing ~50% of ovarian cancer;
- Pol Theta, in solid tumors with genetic mutations in HRD, including BRCA mutations; and
- Werner Helicase, or WRN, in high MSI tumors, present for example, in approximately 15% of colorectal cancer tumors.

Darovasertib (IDE196) / GNAQ/11
Darovasertib (IDE196) is a clinical stage, potent and selective small molecule inhibitor of PKC, a protein kinase that functions downstream of the GTPases GNAQ and GNA11. We hold an exclusive, worldwide license to darovasertib from Novartis, which conducted a Phase 1 clinical trial in metastatic uveal melanoma (MUM), a cancer of the eye with a high frequency of GNAQ or GNA11 gene mutations. This clinical trial demonstrated early clinical activity and tolerability of darovasertib.
IDEAYA is evaluating darovasertib in patients with solid tumors harboring GNAQ or GNA11 mutations. IDEAYA’s clinical development plan for darovasertib in MUM is based on combination therapy, with crizotinib, an investigational cMET inhibitor, in metastatic uveal melanoma (MUM) enabled through our clinical trial collaboration and drug supply agreement with Pfizer. IDEAYA is also evaluating darovasertib and as monotherapy neoadjuvant therapy and adjuvant therapy in primary uveal melanoma (UM).
Activating mutations in GNAQ or GNA11 are found in approximately 90% of uveal melanoma patients, resulting in a dependency on PKC activity which we believe may sensitize these tumors to the effects of darovasertib. Uveal melanoma is a cancer of the eye and the most common primary intraocular malignancy in adults. Approximately 50% of uveal melanoma patients eventually develop metastatic disease following primary treatment, most commonly in the liver.
In addition to uveal melanoma, mutations in GNAQ or GNA11 have also been observed at lower frequencies across other solid tumors, such as cutaneous melanoma and colorectal cancer. GNAQ and GNA11 mutations are included in multiple diagnostic panels.
Prevalence of GNAQ / GNA11 Mutations in Solid Tumors
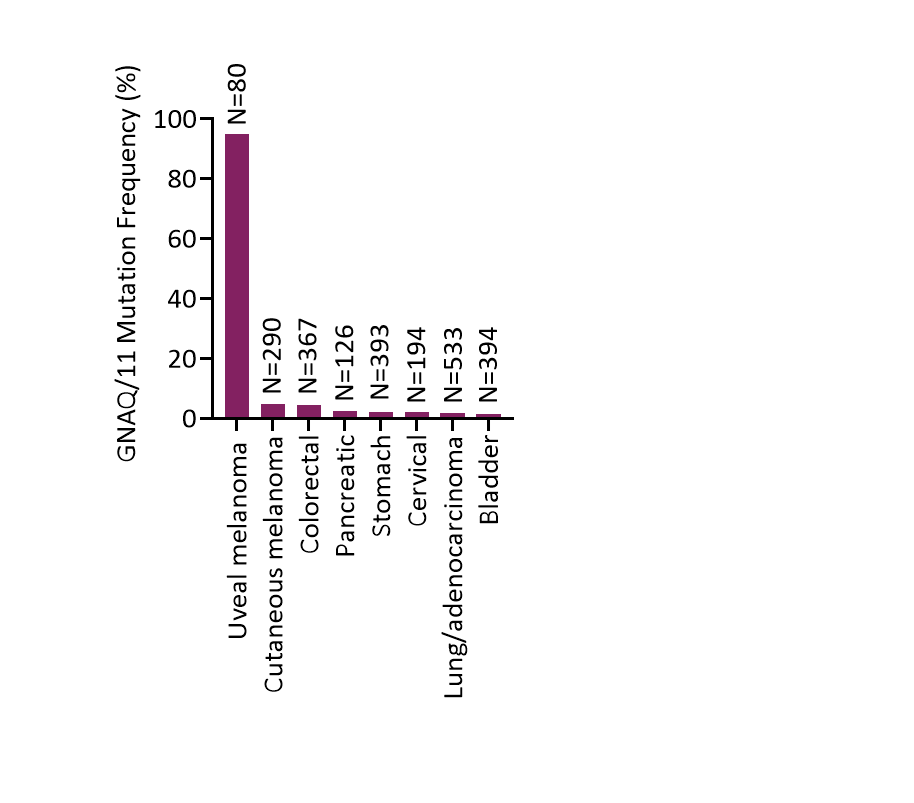
Data from The Cancer Genome Atlas in cBioPortal

MAT2A (IDE397) / MTAP Deletion
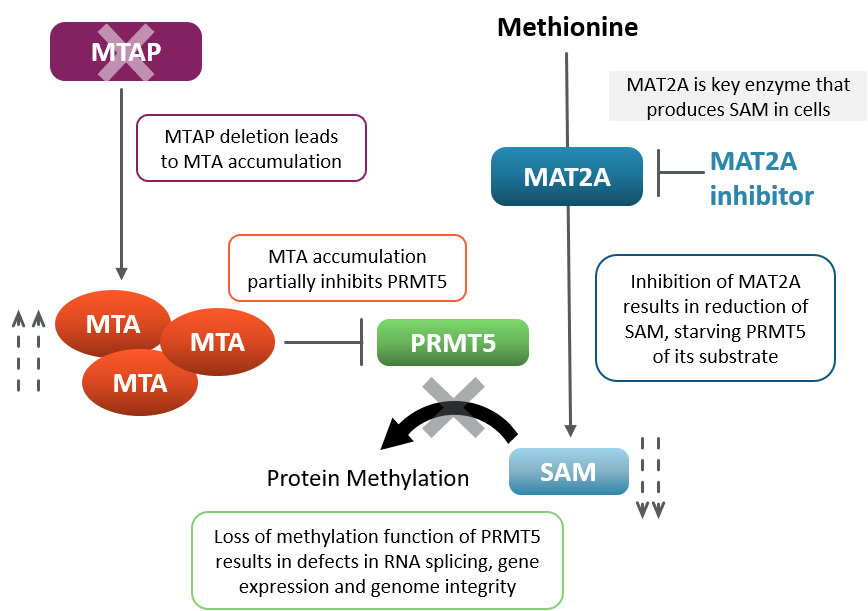
In MTAP null patients, pharmacologically inhibiting MAT2A modulates key protein methylation function, leading to cancer cell death. MTAP-null cells lack the ability to metabolize 5-methylthioadenosine, or MTA, which is an essential step in a biochemical pathway involved in salvaging metabolite S-adenosyl methionine, or SAM. Increased levels of MTA partially inhibit the methyltransferase PRMT5 for which SAM is the substrate. This partial inhibition renders MTAP-null cells more dependent on the activity of methionine adenosyltransferase II alpha or MAT2A, an enzyme that is responsible for the synthesis of SAM. Because of this dependence, loss of MTAP results in synthetic lethality when MAT2A is pharmacologically inhibited.
MTAP deletions are prevalent in approximately 15% of all human tumors across various tumor types. Genetic profiling tests for the deletion of MTAP or for the commonly co-deleted gene CDKN2A are commercially available.
IDEAYA retains all commercial rights to its MAT2A clinical candidate, IDE397.
MTAP-MAT2A Synthetic Lethality Biology
Prevalence of MTAP Deletion in Solid Tumors
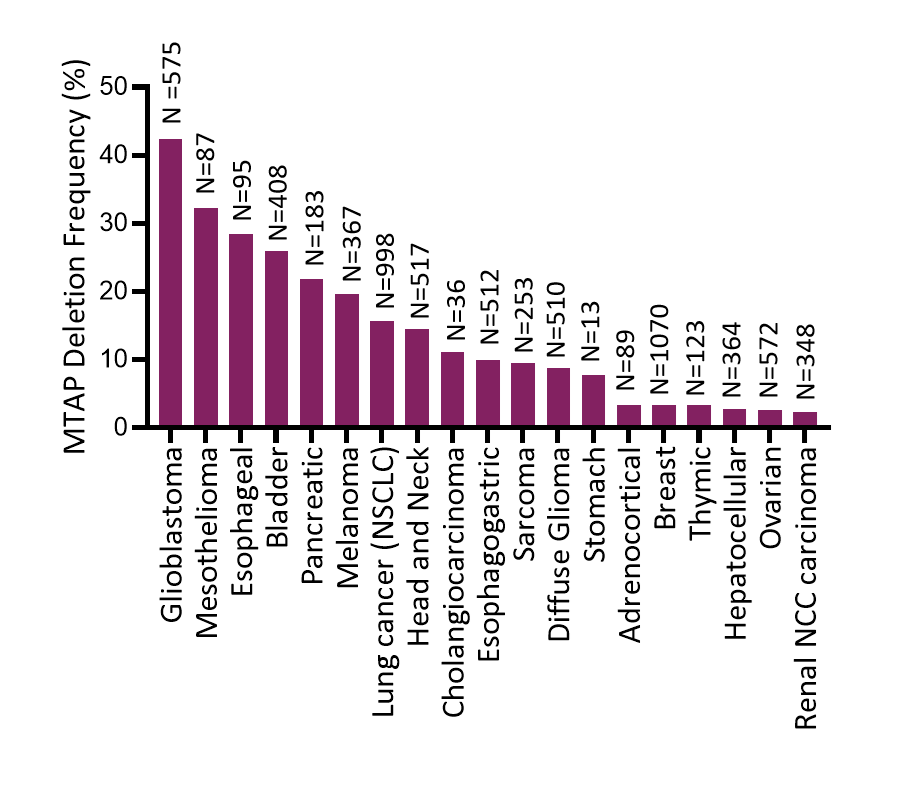
Data from The Cancer Genome Atlas in cBioPortal
DLL3 ADC (IDE849)
DLL3 has been reported to be expressed in multiple solid tumor types, including in SCLC and Neuroendocrine Tumors at approximately 85% and 20-40%, respectively, based on the Human Protein Atlas database. DLL3 has limited extracellular expression in normal tissues, making it a promising therapeutic target in these tumor types, for which there remains significant unmet medical need.
We hold an exclusive license to IDE849, a novel DLL3-targeting Topo-I-payload ADC program, from Jiangsu Hengrui Pharmaceuticals Co., to develop and commercialize IDE849 worldwide outside of Greater China. IDE849 has shown promising antitumor activity in preclinical studies, including tumor regression as a monotherapy in multiple models (IDE849, a novel anti-DLL3 ADC with bystander effect, high DAR and favorable safety profiles. AACR024).
PARG (IDE161) / Defined Biomarker
Poly (ADP-ribose) glycohydrolase, or PARG, has demonstrated a synthetic lethality relationship in cellular and in vivo models with a defined biomarker.
PARG functions as a regulator of DNA repair in the same biochemical pathway as PARP. PARG hydrolyses poly-ADP-ribose (PAR) polymers; pharmacological inhibition of PARG disrupts DNA damage repair cycle and leads to cell death.
IDEAYA retains all commercial rights to its PARG program.
PARG Biology

Pol Theta / Homologous Recombination Deficiency
Pol Theta is involved in a DNA repair process called microhomology mediated end joining, or MMEJ, that is utilized when homologous recombination mediated repair is compromised, as happens in the case of BRCA1 or BRCA2 mutations. The expression of Pol Theta is largely absent in normal cells, but tumor cells harboring double strand break repair defects, such as BRCA1 or BRCA2, show synthetic lethality with Pol Theta inhibition.
Prevalence of HRD Mutations in Solid Tumors

Data from Heeke et al, JCO Precision Oncology, Sep 2018
Werner Helicase / High Microsatellite Instability
MSI is a change in the DNA content of a tumor cell in which the number of repeats of microsatellites, short repeated sequences of DNA, differ as cells divide. High MSI is present in many solid tumor cancers, and tumors are routinely assessed for MSI status in multiple diagnostic profiling tests.
Werner protein is a RecQ family enzyme involved in the maintenance of genome integrity. Germline loss of function mutations in Werner protein lead to premature aging and pre-disposition to cancer. We have evaluated synthetic lethality between Werner protein and high MSI.
Werner protein has two functional domains, and we have shown that the helicase functional domain of Werner protein is responsible for this synthetic lethal interaction in MSI-high cells. We recently published this work in Cell Press – iScience, Werner Syndrome Helicase is Required for the Survival of Cancer Cells with Microsattelite Instability,Vol. 13, pp. 488-497 (Mar 2019).
Prevalence of MSI-High in Solid Tumors

Data from Bonneville et al. JCO Precision Oncology, Sep 2017

